Background¶
Logistic regression is an algorithm that is commonly used for binary classificaiton problems. It does this by predicting a probability and then using a threshold to determine which class to predict. Although the algorithm can be used for multi-class prediction problems, only binary classification is considered in this notebook. The following (sigmoid) function transforms the linear model into the range [0,1]:
$$\sigma(x) = \frac{1}{1+e^{-x}}$$Visualising this function produces the following graph:
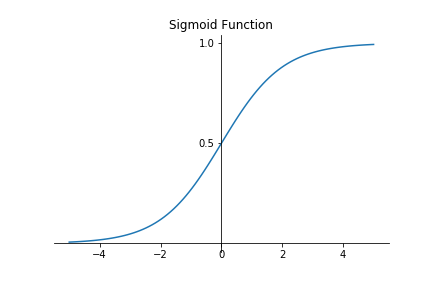
Cost Function¶
The cost function used for linear regression does not work for logistic regression. This is because the sigmoid function causes many local minimums, meaning that finding optimal weights to minimise the cost function proves difficult. Therefore, the cross-entropy loss function is used to produce the following cost function:
$$ \begin{align} J(\theta) = &- \frac{1}{m} \sum^m_{i=1} [y^{(i)} \log(h_{\theta}(x^{(i)})) + (1 - y^{(i)}) \log(1 - h_{\theta}(x^{(i)}))] \\ = &- \frac{1}{m} (y^T \log(h) + (1 - y)^T \log(1-h)) \end{align} $$Where $ h = X \theta$.
Gradient Descent¶
As the logistic regression does not have a closed form solution, like the basic linear regression, gradient descent is used to find the optimal weights that minimise the cost function. The partial derivative of the cost function with respect to each weight is as follows:
$$ \begin{align} \frac{\partial L}{\partial w_j} = [\sigma(X \theta) - y] x_j \end{align} $$This can be used to update the weights as follows:
$$ \begin{align} \theta_{t+1} = \theta_t - \alpha \nabla_{\theta} L \end{align} $$